Advancing Science Through Software
Our goal is to help experimental biologists and bioinformatics scientists manage, organize, validate, and interpret all types of biological data in context, towards experimental validation and testable hypothesis. Our flagship data management, organization solution and bioinformatics expertise can help you on small and large data science projects.
Whether you are interested in characterizing mutational landscape of cancer genomes, understanding perturbed pathways in stem cells, studying trait improvement in agricultural crops, dissecting disease susceptibility/resistance in aquaculture, or performing longitudinal studies to understand microbial dysbiosis in human health, our bioinformatics solutions can help you reduce millions of data points to a manageable and biologically meaningful set of target biomarkers. We follow community accepted best practices for data analysis, supported by rich, interactive visualization.
Microbial Genomics
The development of NGS technologies for assaying microbial genetic material in an environment has become a powerful new approach for rapidly characterizing microbial communities, given a significant percentage of the microbes are uncultivable.
Transcriptomics
RNA-seq with next-generation sequencing (NGS) has become the method of choice for researchers interested in characterizing genome-wide gene and transcript expression in a single experiment.
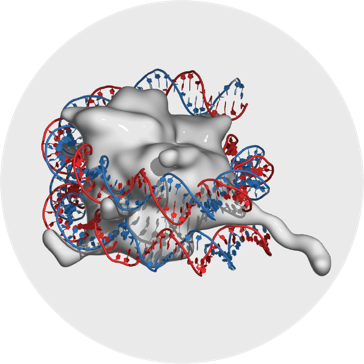
Gene Regulation
Eukaryotic gene regulatory mechanisms that induce or repress the expression of a gene include structural and chemical changes to the chromatin and the DNA, binding of proteins to specific DNA elements, or mechanisms that modulate translation of mRNA.

Variant Discovery
With rapidly falling sequencing costs, whole-exome sequencing (WES) and whole-genome sequencing (WGS) using NGS technologies have become de facto standards to characterize individual genomic landscapes to identify causal genomic variants relevant for understanding disease mechanisms, diagnosis and therapy.
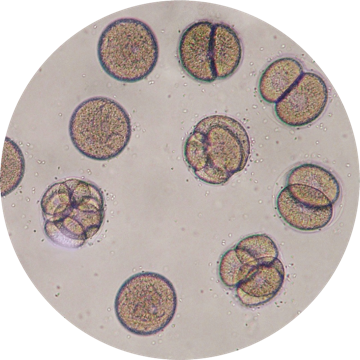
Single-cell Genomics
High-throughput "bulk"-omics techniques have advanced our understanding of the molecular states of biological systems. However, they can only capture ensemble averages of cell states and are poorly suited to understand cell types, states, transitions, and locations. Massively parallel single cell genomics assays that can profile hundreds of thousands of individual cells is rapidly emerging as a revolutionary technology.